The role of epigenetics in vascular disease
Epigenetic changes play an important role in the regulation of gene expression at chromatin level and its deregulation markedly contributes to a plethora of diseases or their aggravation.
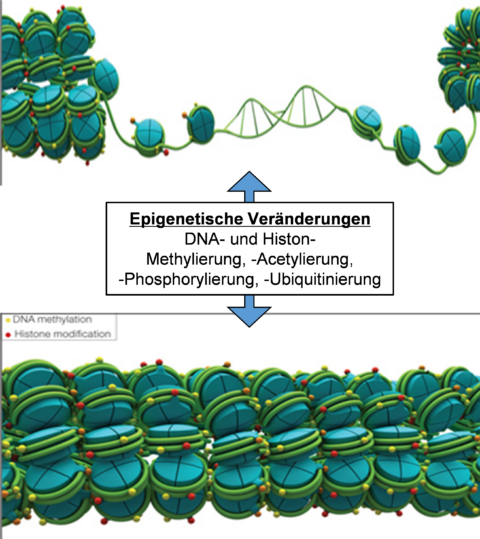
Various modifications of chromatin (open, closed), in particular through histone and DNA methylation, histone acetylation, phosphorylation and ubiquitination, changes the accessibility of transcription factors to the individual genes, preferably their promoter regions. In this manner, gene expression can be significantly affected.
Chronic diseases, atherosclerotic changes within the vessel wall, excessive immunoreaction and other factors can affect the sensitive epigenetic regulation, leading finally to an imbalance in the expression of many genes and thus contributing to the disease progression.
In the case of atherosclerosis or aneurysm, ill changes within the epigenome cause frequently deregulation of the expression of different proteolytic enzymes, augmented degradation of the extracellular matrix (collagen, elastin, proteoglycans) and thus weakening of the vessel wall. These circumstances accelerate the progression of atherosclerosis and the dilation of the aorta, until eventually a rupture occurs, leading finally to a severe clinical outcome.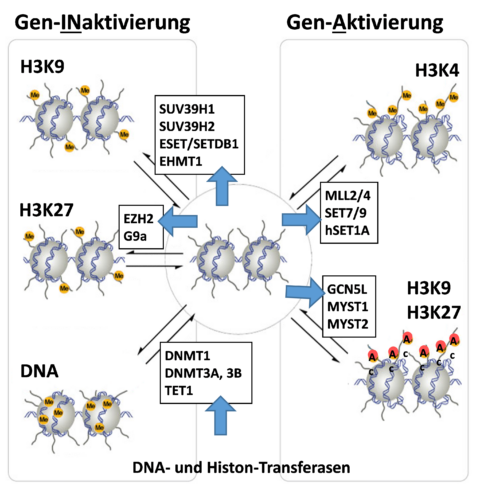
Our results have demonstrated that DNA methylation and histone methylation and acetylation significantly change along with the progression and severity of atherosclerosis [Xu 2019, Xu 2018, Greißel 2016, Culmes 2015, Pelisek 2012]. Such modifications have been found not only in the tissue but also in the blood of the corresponding patients. The same results have been observed also in the advanced abdominal aortic aneurysms (AAA) showing variations in histone methylation and acetylation, as well as in the expressions of the corresponding methyl- and acetyltransferases [Xu 2018, Han 2016].
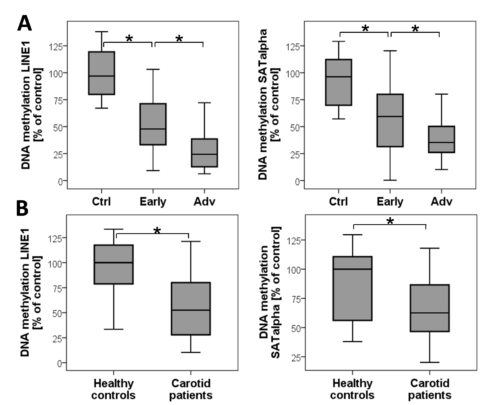
DNA-Methylation in carotid plaques (A) and in blood (B) of the corresponding patients with high-grade carotid stenosis. Quantification was normalised for ALU1. The extent of DNA methylation was normalised for the methylation of DNA in the healthy vessels and set as 100% (Greißel 2016).
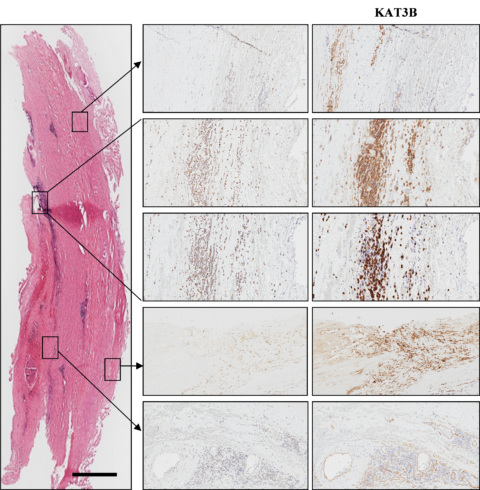
Cellular localisation of histone acetyl transferase KAT3B in AAA (overview picture: Haemalaun-Eosin, scale 1000 µm) (Han 2016).
- Xu S, Kamato D, Little PJ, Nakagawa S, Pelisek J, Jin ZG. Targeting epigenetics and non-coding RNAs in atherosclerosis: from mechanisms to therapeutics. Pharmacol Ther. 2019;196:15-43.
- Xu S, Pelisek J, Jin ZG. Atherosclerosis is an epigenetic disease. Trends Endocrinol Metab. 2018;S1043-2760(18)30090-0.
- Xu S, Xu Y, Yin M, Zhang S, Liu P, Koroleva M, Si S, Little PJ, Pelisek J, Jin ZG. Flow-dependent epigenetic regulation of IGFBP5 by H3K27me3 contributes to endothelial anti-inflammatory effects. Theranostics. 2018;8(11):3007-3021.
- Greißel A, Culmes M, Rudelius M, Burgkart R, Zernecke A, Eckstein HH, Pelisek J. (2015) Histone acetylation and methylation in high graded carotid plaques. Cardiovasc Pathol, 2016;25(2):79-86.
- Han Y, Tanios F, Reeps C, Zhang J, Schwamborn K, Lutz B, Eckstein HH, Zernecke A, Pelisek J. Histone acetylation and expression of corresponding histone acetyl-transferases in human abdominal aortic aneurysm. Clin Epigenetics. 2016; 8:3.
- Culmes M, Greißel A, Napieralski R, Dushenalieva S, Wagner E, Gebhard H, Schmidt M, Eckstein HH, Zernecke A, Pelisek J. DNA methylation in carotid plaques is associated with severity of atherosclerosis and reduced expression of DNA methyltransferases. Thromb Haemost. 2015;21:114(2).
- Yan X, Ehnert S, Culmes M, Claudine C, Schyschka L, Wang Z, Stöckle U, De Sousa PA, Pelisek J, Nussler AK. Small molecule epigenetic inhibitors improve osteogenic differentiation of aged human adipose-derived mesenchymal stem cells by DNA demethylation. PLoS One. 2014;9(6):e90846.
- Culmes M, Eckstein HH, Burgkart R, Nüssler AK, Guenther M, Wagner E, Pelisek J. Endothelial differentiation of Adipose derived mesenchymal stem cells is improved by BIX-01294, an epigenetic modifying drug. Eur J Cell Biol. 2013;92(2):70-79.
- Pelisek J, Eckstein HH, Zernecke A. (2012) Pathophysiological mechanisms of carotid plaque vulnerability: impact on ischemic stroke. Arch Immunol Ther Exp. 2012;60(6):431-442.