Research Group Bojana Müller-Durovic
Our lab investigates how cellular metabolism regulates immune cell function and how metabolic alterations mechanistically contribute to autoimmunity and inflammation, with a focus on Rheumatoid Arthritis and Sjögren’s Disease. By integrating multi-omic data from patient blood and tissue samples, we aim to identify metabolic processes underlying autoimmune pathology. We then use advanced metabolomics techniques and animal models to dissect how these metabolic changes contribute to inflammation. With this approach, we aim to identify novel metabolism-based drug targets for the treatment of rheumatic diseases.
Our Projects
Metabolic rewiring is a key regulator of inflammation. By integrating multi-omics data from patient blood and synovial biopsies, we identified a distinct kynurenine pathway signature in monocytes and macrophages from rheumatoid arthritis patients. This signature closely associates with inflammation in vitro and in mice stimulated with LPS, suggesting that the pathway acts as a mechanism to restrain excessive immune activation. In contrast, in regulatory macrophages the kynurenine pathway is downregulated and remodelled toward NAD de novo synthesis, pointing to divergent regulatory phenotypes of the kynurenine pathway. These data reveal a context-dependent role of the kynurenine pathway in controlling inflammation and offer new opportunities for therapeutic intervention.
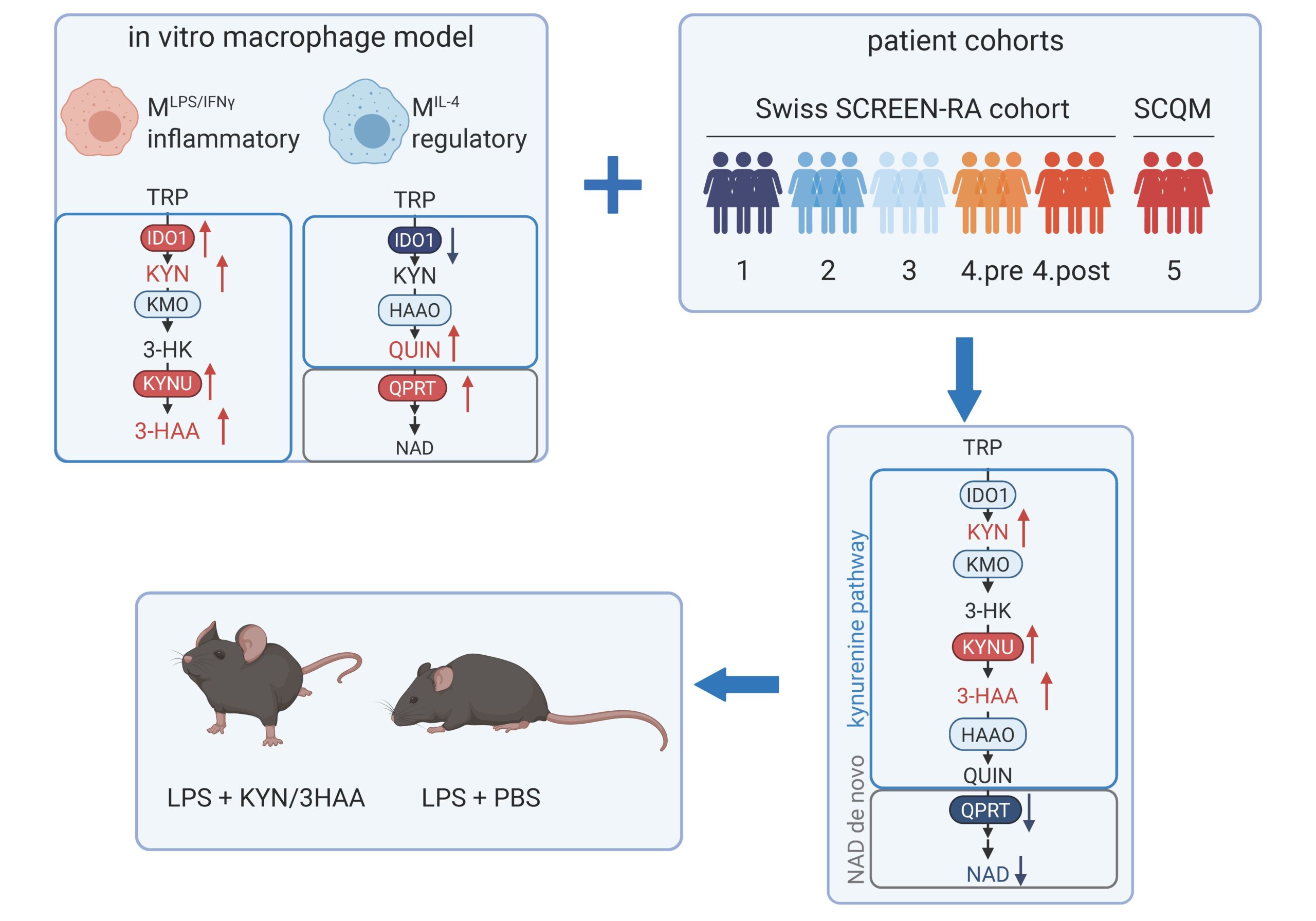
Project lead: Silja Malkewitz
Collaborators: Christian Frezza (CECAD Cologne), Cem Gabay and Matthias Jarlborg (University of Geneva), Axel Finckh (University of Geneva)
Funding: Swiss National Science Foundation, Ambizione grant
Nucleotides are essential not only as building blocks for nucleic acids but also as mediators of signaling and energy balance, while nucleotide sugars serve as key substrates for protein and lipid glycosylation, processes central to immune cell activation and communication. We have observed disruption of these pathways in immune cells of rheumatoid arthritis (RA) patients, which may impact immune cell function and contribute to autoimmunity. Our aim is to dissect the molecular mechanisms underpinning these metabolic dysregulations, elucidate their downstream consequences on immune cell function and regulation, and identify new opportunities for therapeutic intervention in autoimmune diseases.
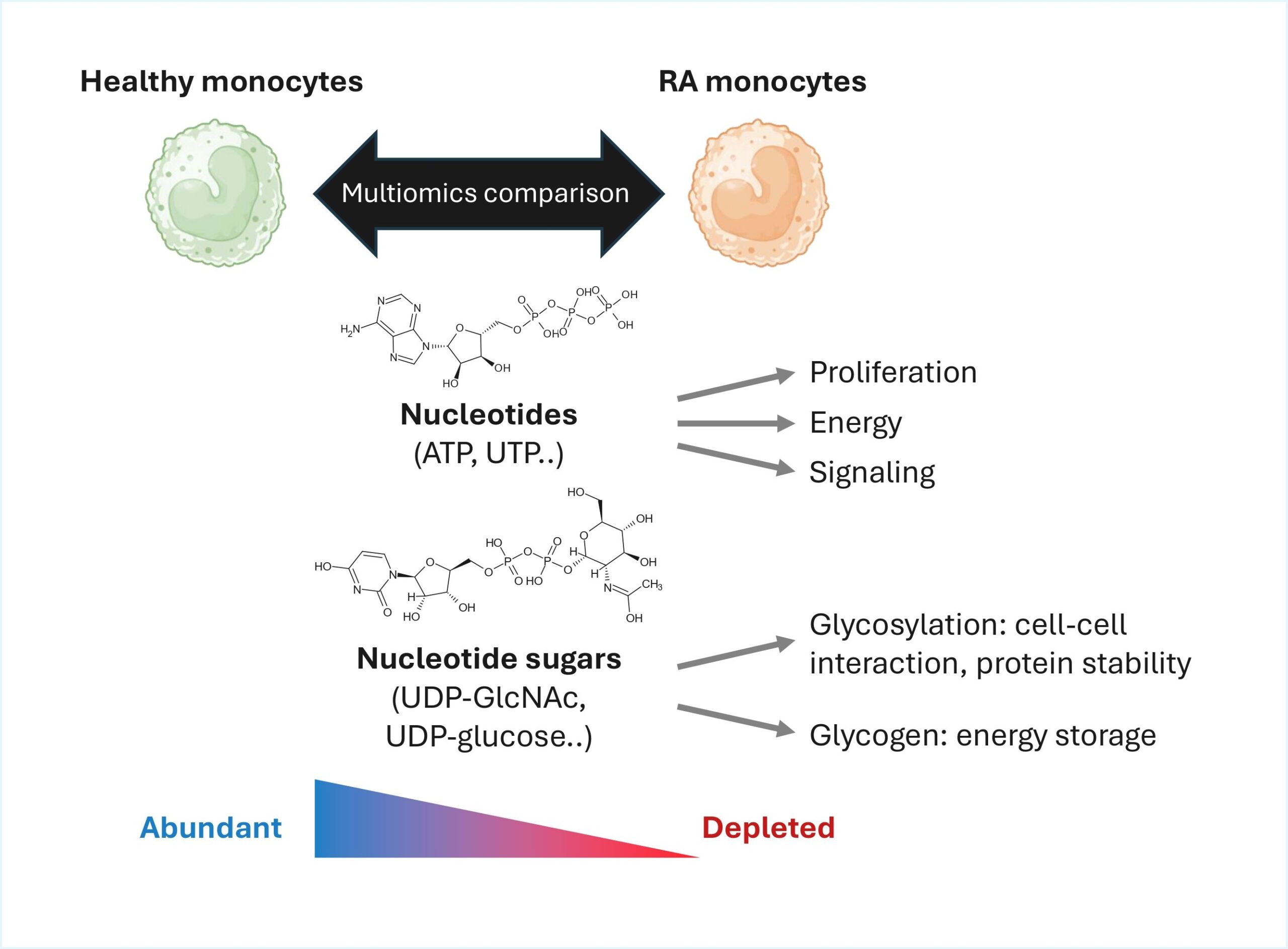
Project lead: Shao Thing Teoh
Funding: UniScientia Foundation
In this project, we are investigating the concept that B cell activation unfolds along a continuum, with lymphoma representing its extreme manifestation, and we aim to understand how cellular metabolism contributes to this process. Sjögren’s Disease provides a powerful model to explore these questions, as it is characterized by pronounced B cell activation and carries one of the highest lymphoma risks among autoimmune conditions. Drawing on the patient clusters defined by Nguyen et al., Lancet Rheumatology 2024, which show that lymphoma develops exclusively in patients with a B cell activation signature in peripheral blood, we will integrate single-cell sequencing with metabolic profiling of B cell subsets from these patients to uncover how metabolism shapes the trajectories of B cells. Our single-cell RNA sequencing analyses of blood B cells from Sjögren’s patients further reveal a striking type I interferon signature accompanied by reduced expression of mitochondrial genes, highlighting a mitochondrial program that we seek to explore as a potential driver of disease pathogenesis.
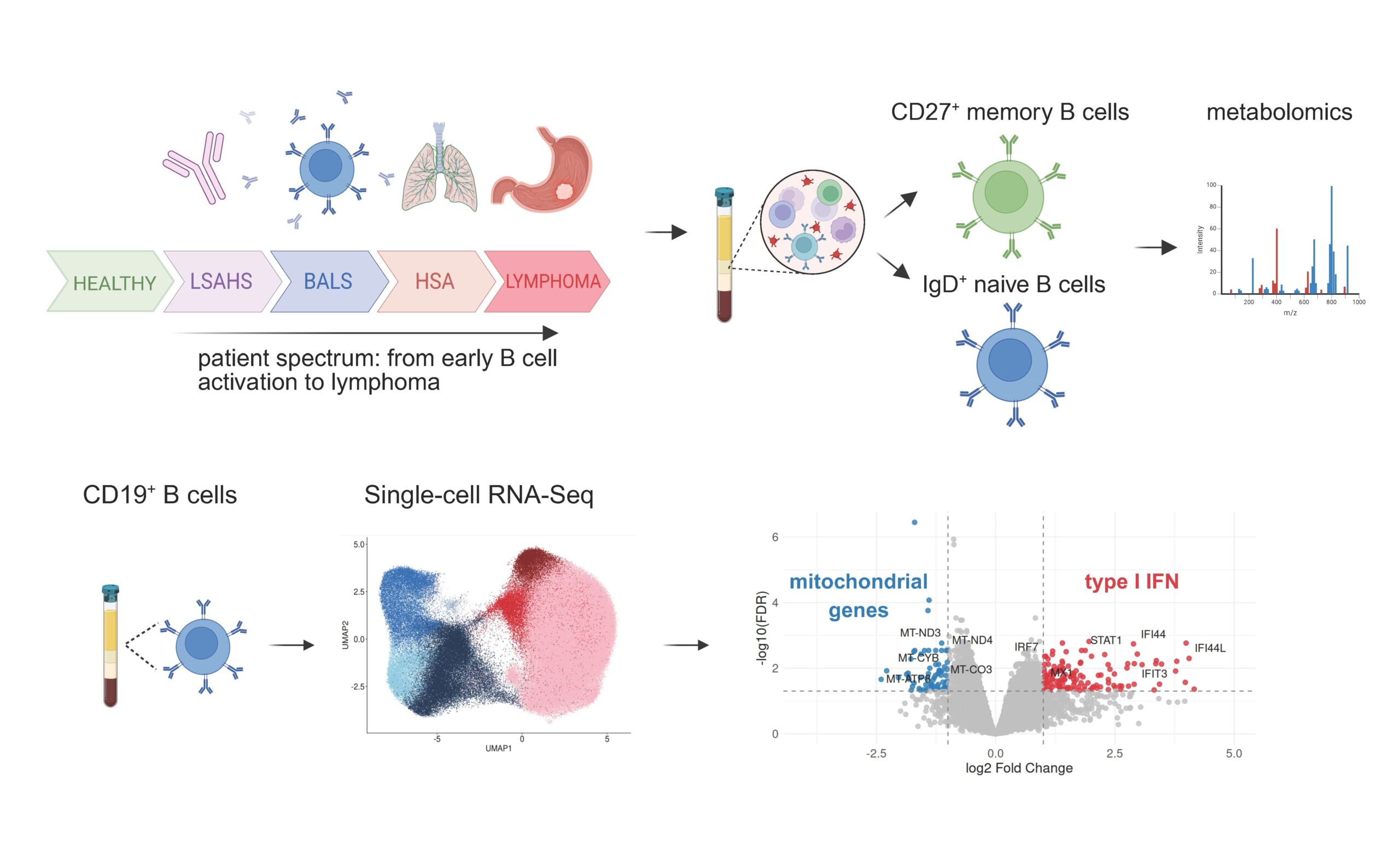
Project lead: Asimina Kakale
Collaborators: Julia Jellusova (Technical University Munich)
Funding: Goldschmidt-Jacobson Foundation, Olga Mayenfisch Foundation
Sjögren’s Disease has a highly heterogeneous presentation, ranging from glandular dryness to serious organ involvement and lymphoma. Current clinical tools cannot reliably predict which patients are at risk of developing systemic complications or lymphoma. Here, we harness high-throughput serum metabolomics and integrate it with blood transcriptomics and rich clinical data from two large French cohorts (ASSESS and Paris-Saclay) to uncover metabolic signatures that define patient subgroups. Using unsupervised multi-omics analysis, we aim to move beyond symptoms to biology, linking metabolic programs with disease activity, trajectories, and lymphoma risk.
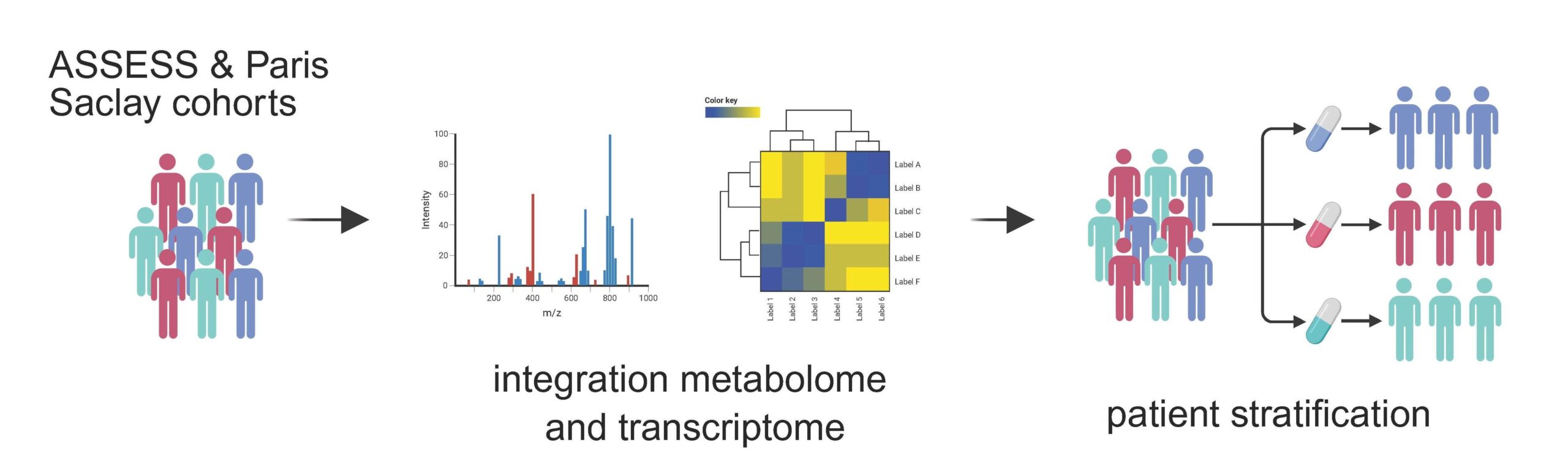
Project lead: Cristian Iperi
Collaborators: Xavier Mariette and Gaetane Nocturne (Université Paris-Saclay)
Funding: Iten-Kohaut Foundation
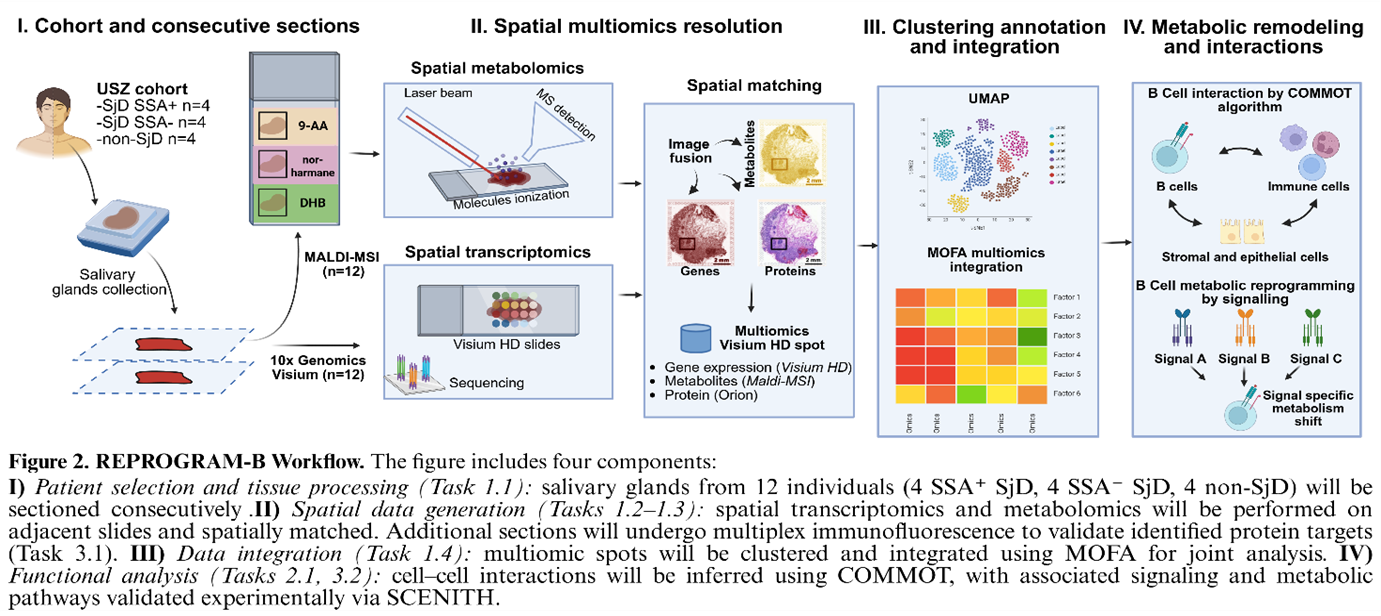
With REPROGRAM-B, we aim to create the first spatially resolved map of B cell metabolism of salivary gland tissue from Sjögren’s Disease patients. By combining spatial metabolomics and spatial transcriptomics with multiplex imaging, we will generate a comprehensive map of how metabolic programs are organized within diseased tissue. Our goal is to identify metabolic vulnerabilities that can be selectively reprogrammed – silencing pathogenic B cells while preserving healthy immunity. This work pioneers spatial multi-omics in autoimmunity.
Project lead: Cristian Iperi
Collaborators: Noel de Miranda and Nina Ogrinc (Leiden University Medical Center), Divi Cornec (LBAI, Université de Bretagne Occidentale)
Funding: Novartis Foundation for Medical-Biological Research
Selected Publications
- Muller-Durovic*, J. Jäger, R. Bantug, C. Hess. Epstein-Barr virus hijacks B cell metabolism to establish persistent infection and drive pathogenesis. Trends Immunol 2024 Jan 2025;46(1):7-16.
https://doi.org/10.1016/j.it.2024.11.011 - Bojana Müller-Durovic, Jessica Jäger, Christine Engelmann, Sabine Altermatt, Yannick Schlup, Nadine Assmann, Urs Duthaler, Patrick Schuhmachers, James West, Lorinda Turner, Leyla Develioglu, Robert Ivanek, Tarik Azzi, Philippe G. Dehio, Jordan Löliger, Marco Geigges, Stefan Dirnhofer, Christian Münz, Glenn R. Bantug, Christoph Hess. A metabolic dependency of EBV can be targeted to hinder B cell transformation. Science, 2024 Jul 5;385(6704).
www.science.org/doi/10.1126/science.adk4898
- Matthias Jarlborg, Post-doc, MD PhD




