Blood
The protein landscape of chronic lymphocytic leukemia (cll)
Meier-Abt F, Lu J, Cannizzaro E, Pohly MF, Kummer S, Pfammatter S, Kunz L, Collins BC, Nadeu F, Lee KS, Xue P, Gwerder M, Roiss M, Hüllein J, Scheinost S, Dietrich S, Campo E, Huber W, Aebersold R, Zenz T. *CCCZ Members in bold
Key points
- The protein landscape of CLL is governed by trisomy 12 and IGHV mutational status.
- Reduced protein abundance buffering implicates the PI3K-AKT-MTOR pathway in the tumorigenic function of trisomy 12.
Many functional consequences of mutations on tumor phenotypes in chronic lymphocytic leukemia (CLL) are unknown. This may be in part due to a scarcity of information on the proteome of CLL. We profiled the proteome of 117 CLL patient samples with data-independent acquisition mass spectrometry (DIA-MS) and integrated the results with genomic, transcriptomic, ex vivo drug response and clinical outcome data. We found trisomy 12, IGHV mutational status, mutated SF3B1, trisomy 19, del(17)(p13), del(11)(q22.3), mutated DDX3X, and MED12 to influence protein expression (FDR < 5%). Trisomy 12 and IGHV status were the major determinants of protein expression variation in CLL as shown by principal component analysis (1055 and 542 differentially expressed proteins, FDR=5%). Gene set enrichment analyses of CLL with trisomy 12 implicated BCR/PI3K/AKT signaling as a tumor driver. These findings were supported by analyses of protein abundance buffering and protein complex formation, which identified limited protein abundance buffering and an upregulated protein complex involved in BCR, AKT, MAPK and PI3K signaling in trisomy 12 CLL. A survey of proteins associated with trisomy 12/IGHV-independent drug response linked STAT2 protein expression with response to kinase inhibitors including BTK and MEK inhibitors. STAT2 was upregulated in U-CLL, trisomy 12 CLL and required for chemokine/cytokine signaling (interferon response). This study highlights the importance of protein abundance data as a non-redundant layer of information in tumor biology, and provides a protein expression reference map for CLL.
Blood. 2021 Jun 29:blood.2020009741. doi: 10.1182/blood.2020009741
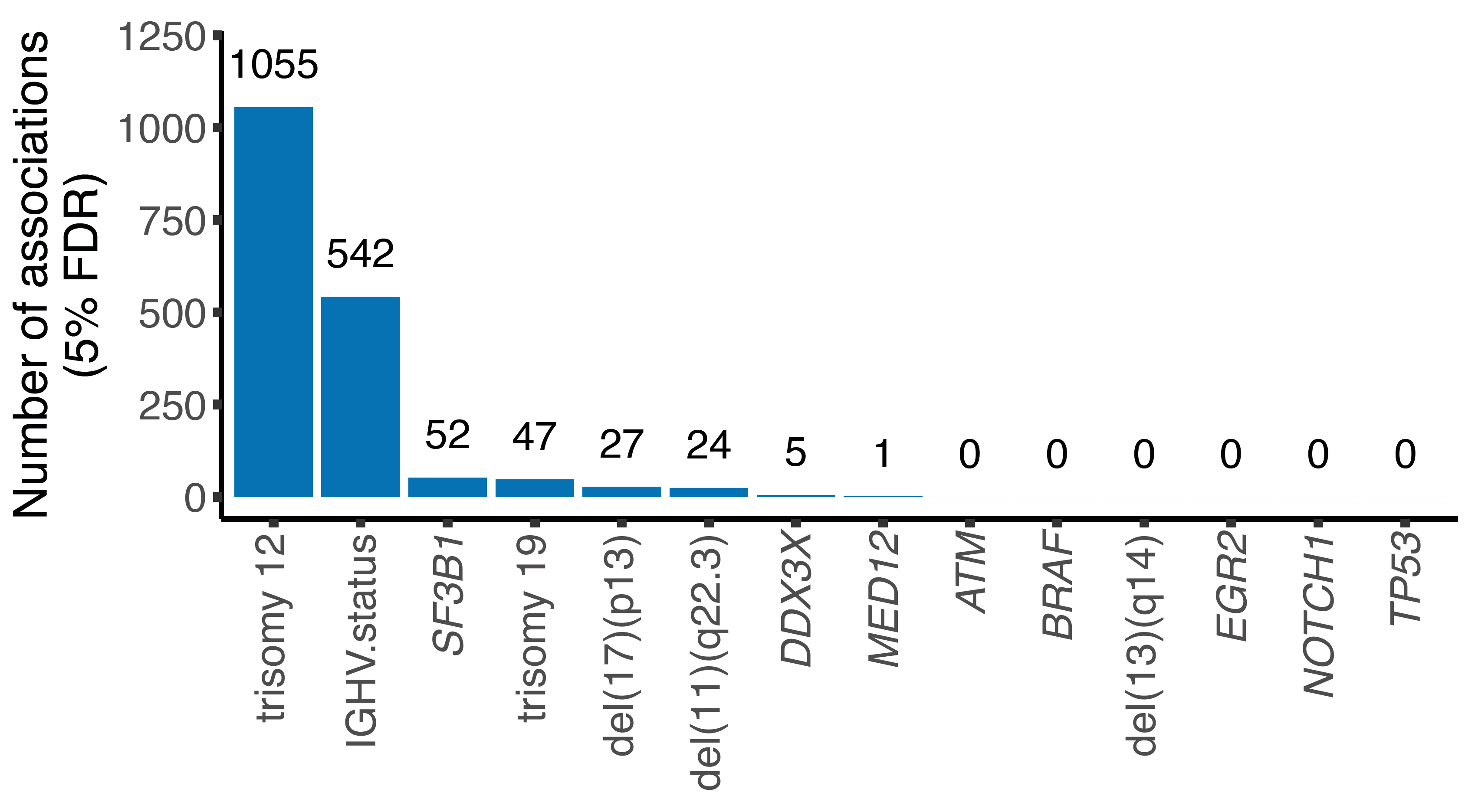
Number of differentially expressed proteins in the main cohort for individual disease drivers in CLL.

